 
You will see various text appear in the console. In the bottom-left tab, labelled "Console", type install.packages("tidyverse") and hit enter. Select the installer appropriate for your operating system You should be able to click "Next" to all dialogs to finish the installation. When the file finishes downloading, double-click to install. 
Where the & enables one to continue using the shell, or close it, without affecting the RStudio instance.Before the first lecture you should have the following software installed on your machine:įollow these instructions if you have not previously installed R, Rstudio or tidyverse, or if you are unsure. Which enables one to then do $ conda activate my_r_env bash_profile so you don't have to type the full path every time, like alias rstudio='/Applications/RStudio.app/Contents/MacOS/RStudio &' Perhaps the only global setting I might recommend is to add an alias for rstudio to your. Instead, one can have many R-based environments and switch between them (or run multiple ones simultaneously) without tampering with global settings. The advantage here is that you aren't fixed to whatever was last set in the environment variables, e.g., via. libPaths() point to the environment-specific locations. Once in R, one can verify that values of R.home() and. Launch RStudio from Activated Conda EnvironmentĪt least for Mac OS X, I find that it is sufficient to activate the environment in a shell session, then launch RStudio. 
(D) Save it as something like run_rstudio.app then just run that and it should work: (C) then delete cat that's already in there and put in:Įxport RSTUDIO_WHICH_R=/Users//anaconda/bin/R (A) open up automator (sorry if you're not on a mac this will only work on mac) (B) then type this in the terminal after it's been sourced (either restart terminal or do source. (A) Put this in your ~/.bash_profile export RSTUDIO_WHICH_R=/Users//anaconda/bin/R (if youre using conda but you could put any R path) Basically, it has to be executed from the correct environment (i.e. 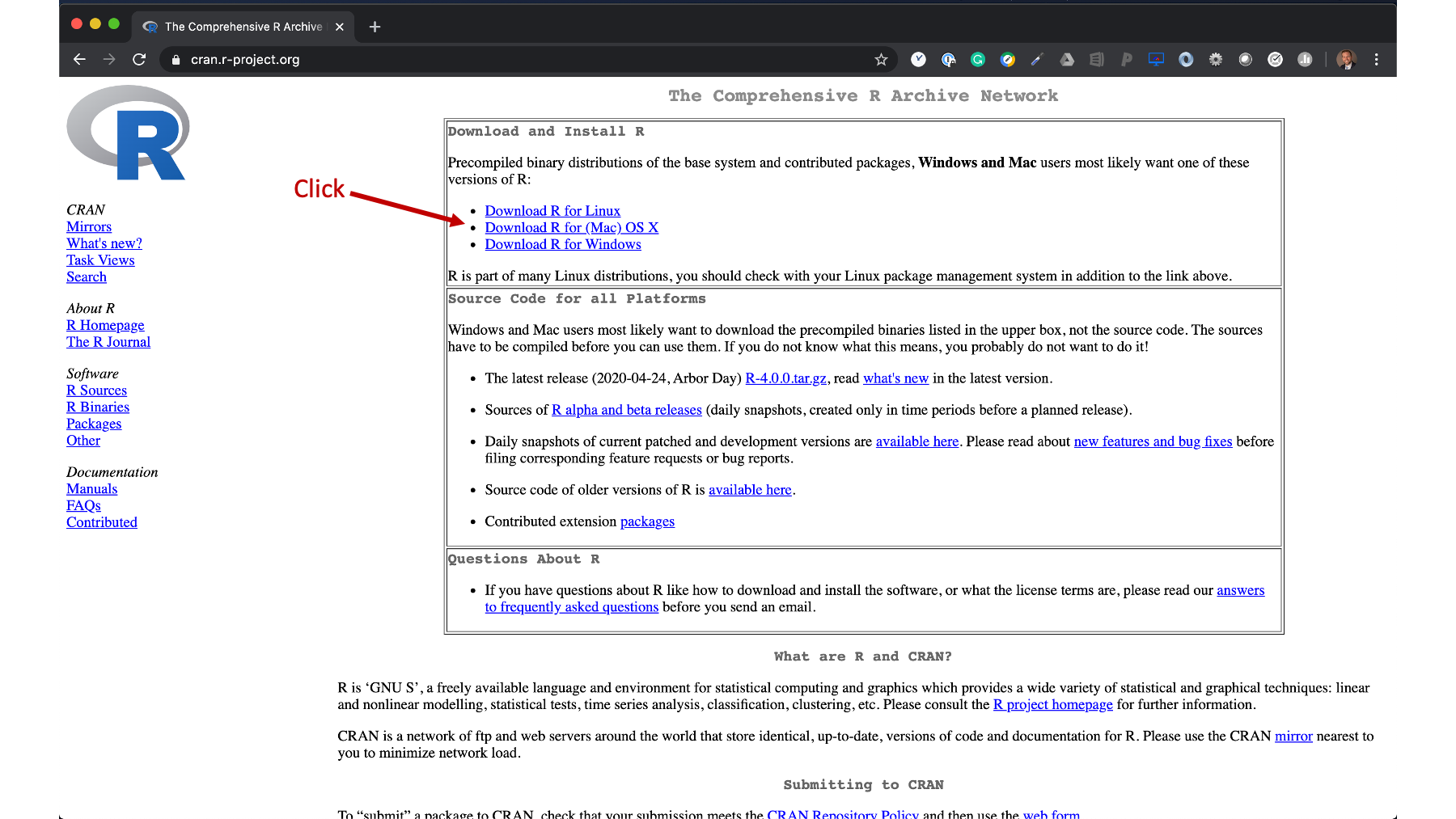
Launchctl setenv RSTUDIO_WHICH_R $RSTUDIO_WHICH_RĪn addition to what Donnelly said above. Update: ADD THIS TO ~/.bash_profile ! export RSTUDIO_WHICH_R="/Users/jespinoz/anaconda/bin/R" For environment variables to be set so that the your desktop environment knows about them, on Linux, you should put them in ~/.profile or else in /etc/pam.d (you may need to logout or shutdown after making those changes) and on OS X, you should check out I think the crux is to launch RStudio from the right environment? Your ~/.bash_profile and ~/.bashrc are only sourced when you run bash. I then closed my terminal and opened a new one (so that the correct conda environment was not activated and successfully launched RStudio with the following command: RSTUDIO_WHICH_R=/home/ray/r_3_3_1-圆4-3.5/bin/R rstudio then force removed the R interpreter ( pacman -Rdd r), then installed r from conda ( conda install -c r r) and it worked fine. So long as which R shows up a working R interpreter (which it should do if you have installed the r package from conda and activated your environment) then launching rstudio from that same environment should pick it up just fine.įor a test, on ArchLinux, I built and installed: Unfortunately, anaconda has no tutorial for this in How to tell RStudio to use R version from Anaconda bashrc but it still didn't work (I sourced both after I added the lines) I tried doing what this guy did and added this to my. When I looked for my R it directed me to: $ which Rīut the directions from (1) is using this path which is very confusing: /Users/jespinoz/anaconda/lib/R/bin/R (2) Launch mac eclipse with environment variables set I'm really not too familiar with environment variables and such things. I've been trying to follow these tutorials but I am lost. bash_profile not sure if this will be useful: $ cat ~/.bash_profileĮxport PATH="/Users/jespinoz/anaconda/bin:$PATH"Įxport RSTUDIO_WHICH_R=/Users/jespinoz/anaconda/bin/R How can I route my conda version of R into RStudio? I want to use RStudio but I don't want to install another version of R. For example, when it outputs errors, it outputs to stdout and splits every character in the string with a linebreak. I ran that version of R (I only installed this version) > version My method of installing R: conda install -c r rĬonda install -channel bioconductor-edger I've been trying to set up my R using conda (eventually to use with Beaker Notebook) and I want to be able to use RStudio with my conda-installed version of R. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
